Cryptomonas
Kerstin Hoef-Emden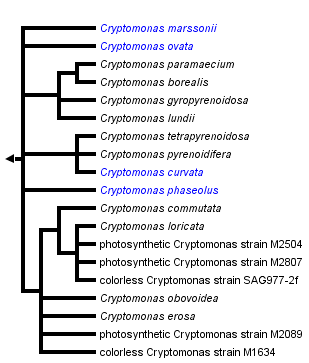

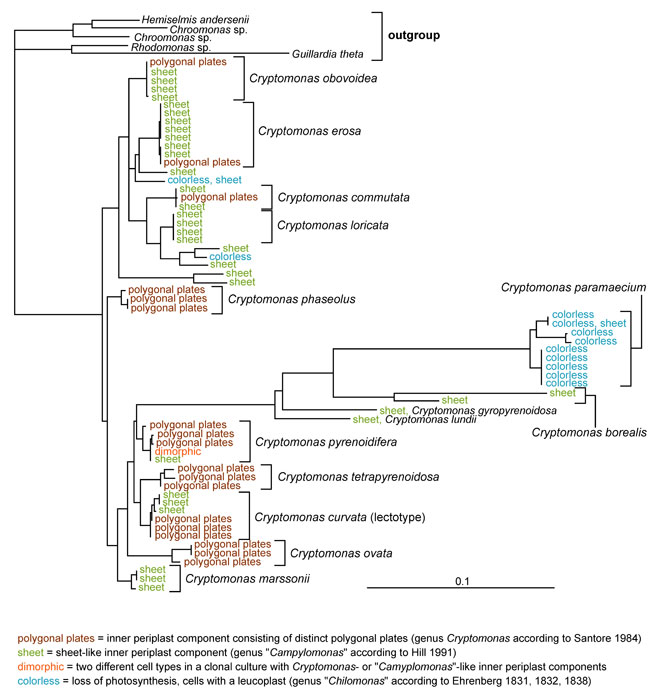
This tree diagram shows the relationships between several groups of organisms.
The root of the current tree connects the organisms featured in this tree to their containing group and the rest of the Tree of Life. The basal branching point in the tree represents the ancestor of the other groups in the tree. This ancestor diversified over time into several descendent subgroups, which are represented as internal nodes and terminal taxa to the right.

You can click on the root to travel down the Tree of Life all the way to the root of all Life, and you can click on the names of descendent subgroups to travel up the Tree of Life all the way to individual species.
For more information on ToL tree formatting, please see Interpreting the Tree or Classification. To learn more about phylogenetic trees, please visit our Phylogenetic Biology pages.
close boxIntroduction
- Described by Ehrenberg (genus: 1831; species: 1832; figures of species: 1838); no type designated
- Lectotype: Cryptomonas curvata (Hoef-Emden and Melkonian 2003)
Characteristics
- Type of biliprotein: PE566 or colorless (leucoplast present; formerly genus Chilomonas; Ehrenberg 1831; Joyon 1963; Sepsenwol 1973)
- Dimorphic
- Light microscopy: olive-green to brownish or colorless freshwater cryptophytes, one deeply bi-lobed plastid or two separate plastids, a cell invagination with a slit-like opening extending into a gullet, with or without pyrenoids, cryptomorphs with straight furrow, campylomorphs with curved furrow and refractive granulum closely appressed to left furrow margin, antapices of campylomorph cells may be bent to the dorsal side
- Ultrastructure of cryptomorphs: inner periplast component made of polygonal plates, surface periplast component consisting of fine fibrils, non-keeled rhizostyle (references summarized in Hill 1991a)
- Ultrastructure of campylomorphs (formerly genus Campylomonas; Hill 1991a): inner periplast component continuous (sheet-like), keeled rhizostyle
- Freshwater only
Discussion of Phylogenetic Relationships
Phylogenetic trees demonstrate that the separation into three genera based only on ultrastructure or pigment content (polygonal periplast plates [Cryptomonas] vs. sheet-like periplast [“Campylomonas”], and phycoerythrin 566 vs. leucoplast [“Chilomonas”]) result in an unnatural system. Instead of forming three clearly separated groups corresponding to morphologically defined genera, strains displaying different morphotypes intermingle in most sub-clades and some strains are even dimorphic with both cell types in a culture (Figure 1; Hoef-Emden and Melkonian 2003; Hoef-Emden 2007). “Chilomonas” not only evolved from photosynthetic species, loss of photosynthesis occurred three times independently (Hoef-Emden 2005). Therefore, the former genera Campylomonas and Chilomonas have been included in Cryptomonas (Hoef-Emden and Melkonian 2003). Until 2003 no valid type, neotype or lectotype of Cryptomonas had been designated. Since the figures by Ehrenberg of Cryptomonas curvata were the only ones that could be clearly assigned by morphology to extant strains, this species has been chosen to be the lectotype species (Hoef-Emden and Melkonian 2003). These examinations also show that Pringsheim's scepticism towards the application of a morphospecies concept in Cryptomonas was justified (Hoef-Emden 2007). Several Cryptomonas species can be identified only by molecular signatures.
References
Ehrenberg CG (1831) Symbolae physicae seu Icones et Descriptiones Animalium Evertebratorum sepositis Insectis que ex itinere per Africanum borealem et Asiam occidentalem Friderici Guilelmi Hemprich et Christiani Godofredi Ehrenberg medicinae et chirurgiae doctorum studio novae aut illustratae redierunt. Mittler, Berlin
Ehrenberg CG (1832) Über die Entwickelung und Lebensdauer der Infusionsthiere; nebst ferneren Beiträgen zu einer Vergleichung ihrer organischen Systeme. Abh. Königl. Akad. Wiss. Berlin, Physik. Klasse 1831: 1-154
Ehrenberg CG (1838) Die Infusionsthiere als vollkommene Organismen: Ein Blick in das tiefere organische Leben der Natur. Nebst einem Atlas von colorirten Kupfertafeln. Voss, Leipzig
Hill DRA (1991a) A revised circumscription of Cryptomonas (Cryptophyceae) based on examination of Australian strains. Phycologia 30: 170-188
Hoef-Emden K (2005) Multiple independent losses of photosynthesis and differing evolutionary rates in the genus Cryptomonas (Cryptophyceae): Combined phylogenetic analyses of DNA sequences of the nuclear and nucleomorph ribosomal operons. J. Mol. Evol. 60: 183-195
Hoef-Emden K (2007) Revision of the genus Cryptomonas (Cryptophyceae) II: Incongruences between classical morphospecies and molecular phylogeny in smaller pyrenoid-less cells. Phycologia 46: 402-428
Hoef-Emden K, Melkonian M (2003) Revision of the genus Cryptomonas (Cryptophyceae): a combination of molecular phylogeny and morphology provides insights into a long-hidden dimorphism. Protist 154: 371-409
Joyon L (1963) Sur la présence d'un leucoplaste chez la Cryptomonadine decolorée Chilomonas paramecium (Ehrenberg). C. R. Hebd. Séances Acad. Sci. 256: 3502-3503
Sepsenwol S (1973) Leucoplast of the cryptomonad Chilomonas paramecium. Exp. Cell Res. 76: 395-409
Skuja H (1948) Taxonomie des Phytoplanktons einiger Seen in Uppland, Schweden. Symb. Bot. Ups. 9: 346-367
Title Illustrations

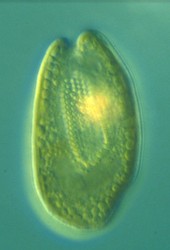
| Scientific Name | Cryptomonas sp. |
|---|---|
| Comments | from a freshwater sample. Cryptomonas lundii/borealis relationship. |
| Image Use |
 This media file is licensed under the Creative Commons Attribution-NonCommercial License - Version 3.0. This media file is licensed under the Creative Commons Attribution-NonCommercial License - Version 3.0.
|
| Copyright |
© 2008 Kerstin Hoef-Emden

|
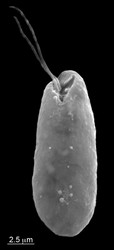
| Scientific Name | Cryptomonas paramaecium CCAP977/2a |
|---|---|
| Specimen Condition | Dead Specimen |
| Image Use |
 This media file is licensed under the Creative Commons Attribution-NonCommercial License - Version 3.0. This media file is licensed under the Creative Commons Attribution-NonCommercial License - Version 3.0.
|
| Copyright | © 2008 Natalie Donaher and John Archibald |
About This Page
This page is being developed as part of the Tree of Life Web Project Protist Diversity Workshop, co-sponsored by the Canadian Institute for Advanced Research (CIFAR) program in Integrated Microbial Biodiversity and the Tula Foundation.
Kerstin Hoef-Emden

Universität zu Köln, Köln, Germany
Correspondence regarding this page should be directed to Kerstin Hoef-Emden at
Page copyright © 2010 Kerstin Hoef-Emden
 Page: Tree of Life
Cryptomonas .
Authored by
Kerstin Hoef-Emden.
The TEXT of this page is licensed under the
Creative Commons Attribution-NonCommercial License - Version 3.0. Note that images and other media
featured on this page are each governed by their own license, and they may or may not be available
for reuse. Click on an image or a media link to access the media data window, which provides the
relevant licensing information. For the general terms and conditions of ToL material reuse and
redistribution, please see the Tree of Life Copyright
Policies.
Page: Tree of Life
Cryptomonas .
Authored by
Kerstin Hoef-Emden.
The TEXT of this page is licensed under the
Creative Commons Attribution-NonCommercial License - Version 3.0. Note that images and other media
featured on this page are each governed by their own license, and they may or may not be available
for reuse. Click on an image or a media link to access the media data window, which provides the
relevant licensing information. For the general terms and conditions of ToL material reuse and
redistribution, please see the Tree of Life Copyright
Policies.
- First online 14 September 2008
- Content changed 02 April 2010
Citing this page:
Hoef-Emden, Kerstin. 2010. Cryptomonas . Version 02 April 2010 (under construction). http://tolweb.org/Cryptomonas/97214/2010.04.02 in The Tree of Life Web Project, http://tolweb.org/






 Go to quick links
Go to quick search
Go to navigation for this section of the ToL site
Go to detailed links for the ToL site
Go to quick links
Go to quick search
Go to navigation for this section of the ToL site
Go to detailed links for the ToL site